Plotting empirical logits in R
| Linear regression models assume that the conditional expectation of the dependent variable, E[y | x], is linear in the predictor variables x (modulo any polynomial terms). Simple scatterplots are often useful to check if linearity appears reasonable, or if it requires applying some transformation to a predictor variable. |
Analogously, logit models assume that the logit-transformed response probability (i.e. the log odds of response) is linear in the predictors. Empirical logit plots are a straightforward analogue of scatterplots for checking this assumption. Since our response variable is binary, we can’t directly logit-transform the variable. But we can place the binary values in equally-sized bins, estimate “local” probabilities by averaging within each bin, and then logit-transform those probabilities. These are our empirical logits, which we can then plot.
The following R function does all this:
emplogit = function(x, y, binsize = NULL, ci = FALSE, probit = FALSE,
prob = FALSE, main = NULL, xlab = "", ylab = ""){
# x vector with values of the independent variable
# y vector of binary responses
# binsize integer value specifying bin size (optional)
# ci logical value indicating whether to plot approximate
# confidence intervals (not supported as of 02/08/2015)
# probit logical value indicating whether to plot probits instead
# of logits
# prob logical value indicating whether to plot probabilities
# without transforming
#
# the rest are the familiar plotting options
if (length(x) != length(y))
stop("x and y lengths differ")
if (any(y < 0 | y > 1))
stop("y not between 0 and 1")
if (length(x) < 100 & is.null(binsize))
stop("Less than 100 observations: specify binsize manually")
if (is.null(binsize)) binsize = min(round(length(x)/10), 50)
if (probit){
link = qnorm
if (is.null(main)) main = "Empirical probits"
} else {
link = function(x) log(x/(1-x))
if (is.null(main)) main = "Empirical logits"
}
sort = order(x)
x = x[sort]
y = y[sort]
a = seq(1, length(x), by=binsize)
b = c(a[-1] - 1, length(x))
prob = xmean = ns = rep(0, length(a)) # ns is for CIs
for (i in 1:length(a)){
range = (a[i]):(b[i])
prob[i] = mean(y[range])
xmean[i] = mean(x[range])
ns[i] = b[i] - a[i] + 1 # for CI
}
extreme = (prob == 1 | prob == 0)
prob[prob == 0] = min(prob[!extreme])
prob[prob == 1] = max(prob[!extreme])
g = link(prob) # logits (or probits if probit == TRUE)
linear.fit = lm(g[!extreme] ~ xmean[!extreme])
b0 = linear.fit$coef[1]
b1 = linear.fit$coef[2]
loess.fit = loess(g[!extreme] ~ xmean[!extreme])
plot(xmean, g, main=main, xlab=xlab, ylab=ylab)
abline(b0,b1)
lines(loess.fit$x, loess.fit$fitted, lwd=2, lty=2)
}
Now for an example. Suppose that the population model is:
\(\text{logit}(p_i) = \beta_0 + \beta_1 \text{log}(x)\) \(y_i \sim \text{Bernoulli}(p_i)\)
We generate a sample that conforms to this model:
set.seed(1234)
n = 2000
x1 = exp(rnorm(n/2, mean=0, sd=1))
x2 = exp(rnorm(n/2, mean=1, sd=1))
x = c(x1, x2)
y = c(rep(0, n/2), rep(1, n/2))
If we aren’t sure about the true population model, we might not know which of \(x\) or \(\text{log}(x)\) to use in a logit model of \(y\). Looking at the empirical logits makes the choice obvious:
emplogit(x, y)
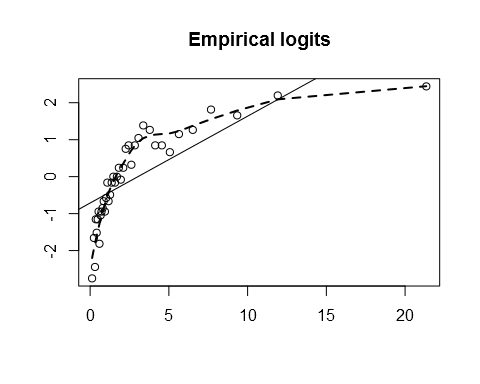
emplogit(log(x), y)
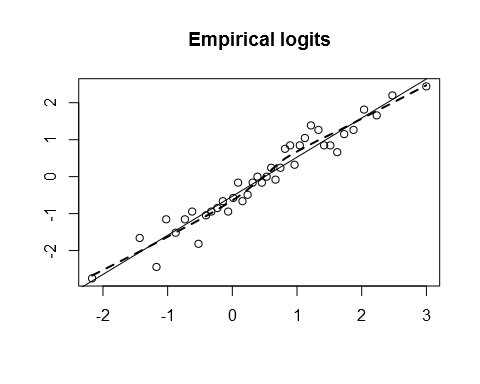
In this artificial example, the choice of transformation makes a big difference to the interpretation of the model. Let’s fit the two models:
m1 = glm(y ~ x, family="binomial")
m2 = glm(y ~ log(x), family="binomial")
The misspecified model m1 gives us an estimate \(\hat{\beta_1}\) = 0.38, for an estimated marginal probability effect of 0.09 (at \(p_i\) = 0.5). The correctly specified model m2 outputs \(\hat{\beta_1}\) = 1.06, and the estimated marginal probability effect is 0.53 (at \(p_i\) = 0.5 and the corresponding geometric mean of \(x\), 1.64).